
|
ROCPLOT documentation
|
CONTENTS
1.0 SUMMARY
2.0 INPUTS & OUTPUTS
3.0 INPUT FILE FORMAT
4.0 OUTPUT FILE FORMAT
5.0 DATA FILES
6.0 USAGE
7.0 KNOWN BUGS & WARNINGS
8.0 NOTES
9.0 DESCRIPTION
10.0 ALGORITHM
11.0 RELATED APPLICATIONS
12.0 DIAGNOSTIC ERROR MESSAGES
13.0 AUTHORS
14.0 REFERENCES
1.0 SUMMARY
Provides interpretation and graphical display of the performance of discriminating elements (e.g. profiles for protein families). rocplot reads file(s) of hits from discriminator-database search(es), performs ROC analysis on the hits, and writes graphs illustrating the diagnostic performance of the discriminating elements.
Perform ROC analysis on hits files
2.0 INPUTS & OUTPUTS
ROCPLOT reads a directory of one or more hits files and writes a text,
summary file containing ROC value(s), which are a convenient numerical
measure of the sensitivity and specificity of a predictive method. GNUPLOT
files for the following graphs are also written.
(i) ROC plots displaying graphically the method sensitivity and specificity.
(ii) Classification plots, which are a useful aid in interpreting ROC plots
and ROC values.
(iii) In some modes (see below) a bar chart of the distribution of ROC values
is generated.
2.1 ROCPLOT modes
ROCPLOT runs in one of two basic modes:
(i) "Single hits file"
(ii) "Multiple hits file".
2.1.1 Single hits file mode
ROC analysis is performed on the single hits file. A ROC plot containing
one ROC curve and a single ROC value and classification plot are generated.
2.1.2 Multiple hits files mode
The same ROC number must be given in the hits files and each file must
contain at least this number of non-TRUE hits (see Section 3.1): an error
is generated and the program terminates otherwise.
In "multiple hits file mode" there are two sub-modes:
(i) "Do not combine data"
(ii) "Combine data".
2.1.3 Do not combine data mode
ROC analysis is performed separately for each hits file. Multiple ROC curves
are given on the same ROC plot. A ROC value and classification plot are
generated for each hits file. A bar chart giving the distribution of ROCn
values is also generated. The mean and standard deviation of ROCn values are
written to the summary file.
2.1.4 Combine data mode
The hits are combined and ROC analysis is performed on the whole (see Section
9.6). A ROC plot containing one ROC curve and a single ROC value and
classification plot are generated.
In "combine data" mode there are a further two sub-modes:
(i) "Single gold standard"
(ii) "Multiple gold standard".
These determine how the ROC number and value are calculated.
2.1.5 Single gold standard mode
There is a single gold standard (list of known true hits) for the different
searches. The same number of known true hits must be specified in the hits
files: an error is generated and the program terminates otherwise. The
accession number (or other code) and start and end point of each hit must
also be given (see Section 3.1).
2.1.6 Multiple gold standard mode
There is a gold standard for each different search.
The output in the different modes is summarised (Figure 1).
Figure 1 Summary of ROCPLOT output
____________________________________________________
| SINGLE HITS FILE | MULTIPLE HITS FILES |
| | | |
| | Do not combine | Combine |
| | data | data |
_____________________|___________________|________________|_____________|
| | | |
ROC curves / value | Single | Multiple (1) | Single |
Bar chart | - | Yes | - |
Classification plot | Single | Multiple | Single |
Summary file | Yes | Yes | Yes |
_____________________|___________________|________________|_____________|
|
(1) Multiple ROC curves are given on a single ROC plot.
3.0 INPUT FILE FORMAT
3.1 Hits files
A hits file contains a list of classified hits that are
rank-ordered on the basis of score. The first line must have '>' in the
first character position and a space (' ') in the second, then two token
- integer pairs delimited by ';'. The integer following 'RELATED' is the
total number of known true hits ('relatives') and is the maximum number of
TRUE tokens (see below) that could ever appear in the hits file. The
integer following 'ROC' is the ROC value that will be calculated. This
integer also determines the limit of the x-axes of the ROC and classification
plots (see Sections 9.2 & 9.4).
The file then contains a number of lines corresponding to a list of
classified hits. The hits *must* be rank-ordered on the basis of score,
p-value, E-value etc, with the highest scoring / most significant hit given
in the highest rank (1); i.e. on the second line of the file. Other hits
should then be given in order of decreasing score / significance.
The first string in a hit line is the classification and must be one of the
following: 'TRUE', 'CROSS', 'UNCERTAIN', 'UNKNOWN' or 'FALSE'. If ROCPLOT
is run in "Multiple hits files" - "Combine data" - "Single gold standard"
modes, each hit line must contain a second string followed by 2 integers.
These are required so that ROCPLOT can identify unique hits in the lists of
hits (see Section 10.4). For hits to sequences, the string is the accession
number (or other database code) and the integers are the start and end point
of the hit relative to the full length sequence. For some applications the
start and end point data are not required to define unique hits: in these
cases the start and end values for all hits should be set to 0 and 1
respectively.
4.0 OUTPUT FILE FORMAT
4.0 OUTPUT FILE FORMAT
4.1 Summary file
The summary file is shown in Figure 3. The first section is comments including
the modes ROCPLOT was run in. The file may then contain a section where the
file name, number of known true hits and ROCn value are given for each hits
file. In cases where data from multiple hits files were combined a single
ROCn value will be given instead of this section. The mean and SD of the ROCn
values are given if calculated.
4.2 GNUPLOT files
ROCPLOT generates various gnuplot driver and data files depending upon mode.
For example, the user specifies the base name of the rocplot, classification,
bar chart and summary files to be "_rocplot", "_classplot", "_barchart" and
"_summary" respectively. If ROCPLOT is run in "Multiple hits files" -
"Combine data" - "Single gold standard" mode the following files are
generated.
_classplot_dat0 Data file for classification plot
_classplot_dat1 Data file for classification plot
_classplot_dat2 Data file for classification plot
_classplot_dat3 Data file for classification plot
_classplot_dat4 Data file for classification plot
_classplot Driver file for classification plot
_rocplot_dat0 Data file for roc plot.
_rocplot Driver file for roc plot.
_summary Summary file.
|
If ROCPLOT is run in "Multiple hits files" - "Combine data" - "Single gold
standard" mode the following files are generated.
_classplot0_dat0 Data file for first classification plot
_classplot0_dat1 ""
_classplot0_dat2 ""
_classplot0_dat3 ""
_classplot0_dat4 ""
_classplot0 Driver file for first classification plot
_classplot1_dat0 Data file for second classification plot
_classplot1_dat1 ""
_classplot1_dat3 ""
_classplot1_dat4 ""
_classplot1 Driver file for second classification plot
_rocplot_dat0 Data file for roc plot.
_rocplot_dat1 ""
_rocplot Driver file for roc plot.
_summary Summary file.
|
Note that there is no _classplot1_dat2 indicating that the second hits file
did not contain any hits for one of the data series (see Section 9.4).
If ROCPLOT is run in "Multiple hits files" - "Do not combine data" the
following files are generated.
_barchart_dat Data file for bar chart.
_barchart Driver file for bar chart.
|
The plots are visualised by using GNUPLOT, for example by typing load
'_classplot1' from the GNUPLOT command line.
Output files for usage example
File: _rocplot
# GNUPLOT driver file for roc plot
set title "ROC plots for data1.hits & data2.hits (combined - "
set xlabel "1 - SPEC"
set ylabel "SENS"
set nokey
set noautoscale
set xrange [0:1]
set yrange [0:1]
set key top outside title "Data Series" box 3
set data style points
set pointsize 0.45
plot "_rocplot_dat0" smooth bezier title "Combined dataset (0.185)"
|
File: _rocplot_dat0
# GNUPLOT data file for rocplot, series 0
0.000 0.007
0.000 0.014
0.000 0.021
0.000 0.029
0.200 0.029
0.167 0.036
0.143 0.043
0.250 0.043
0.222 0.050
0.200 0.057
0.182 0.064
0.250 0.064
0.231 0.071
0.214 0.079
0.200 0.086
0.250 0.086
0.235 0.093
0.222 0.100
0.263 0.100
0.250 0.107
0.238 0.114
0.273 0.114
0.261 0.121
0.292 0.121
0.280 0.129
0.308 0.129
0.296 0.136
0.286 0.143
0.276 0.150
0.300 0.150
0.290 0.157
0.281 0.164
0.273 0.171
0.294 0.171
0.286 0.179
0.278 0.186
0.297 0.186
0.316 0.186
0.333 0.186
0.350 0.186
0.366 0.186
0.381 0.186
0.395 0.186
0.409 0.186
0.400 0.193
0.391 0.200
0.404 0.200
0.417 0.200
0.429 0.200
0.440 0.200
0.451 0.200
0.462 0.200
0.472 0.200
0.481 0.200
0.473 0.207
0.464 0.214
0.474 0.214
0.483 0.214
0.492 0.214
0.500 0.214
0.508 0.214
0.516 0.214
0.524 0.214
0.531 0.214
0.538 0.214
0.545 0.214
0.552 0.214
0.559 0.214
0.565 0.214
0.571 0.214
0.577 0.214
0.583 0.214
0.589 0.214
0.595 0.214
0.600 0.214
0.605 0.214
0.610 0.214
0.615 0.214
0.620 0.214
0.625 0.214
|
File: _classplot
# GNUPLOT driver file for classification plot
set title "Classification plot for data1.hits & data2.hits (c"
set xlabel "Number of hits detected"
set ylabel "Proportion of hits detected that are of a certain type"
set nokey
set key top outside title "Data Series" box 3
set data style points
set pointsize 0.45
plot "_classplot_dat0" smooth bezier title "True hits", "_classplot_dat1" smooth bezier title "Cross hits", "_classplot_dat2" smooth bezier title "Uncertain hits", "_classplot_dat3" smooth bezier title "Unknown hits", "_classplot_dat4" smooth bezier title "False hits"
|
File: _classplot_dat0
# GNUPLOT data file for True hits, series 0
1.000 1.000
2.000 1.000
3.000 1.000
4.000 1.000
5.000 0.800
6.000 0.833
7.000 0.857
8.000 0.750
9.000 0.778
10.000 0.800
11.000 0.818
12.000 0.750
13.000 0.769
14.000 0.786
15.000 0.800
16.000 0.750
17.000 0.765
18.000 0.778
19.000 0.737
20.000 0.750
21.000 0.762
22.000 0.727
23.000 0.739
24.000 0.708
25.000 0.720
26.000 0.692
27.000 0.704
28.000 0.714
29.000 0.724
30.000 0.700
31.000 0.710
32.000 0.719
33.000 0.727
34.000 0.706
35.000 0.714
36.000 0.722
37.000 0.703
38.000 0.684
39.000 0.667
40.000 0.650
41.000 0.634
42.000 0.619
43.000 0.605
44.000 0.591
45.000 0.600
46.000 0.609
47.000 0.596
48.000 0.583
49.000 0.571
50.000 0.560
51.000 0.549
52.000 0.538
53.000 0.528
54.000 0.519
55.000 0.527
56.000 0.536
57.000 0.526
58.000 0.517
59.000 0.508
60.000 0.500
61.000 0.492
62.000 0.484
63.000 0.476
64.000 0.469
65.000 0.462
66.000 0.455
67.000 0.448
68.000 0.441
69.000 0.435
70.000 0.429
71.000 0.423
72.000 0.417
73.000 0.411
74.000 0.405
75.000 0.400
76.000 0.395
77.000 0.390
78.000 0.385
79.000 0.380
80.000 0.375
|
File: _classplot_dat1
# GNUPLOT data file for Cross hits, series 1
1.000 0.000
2.000 0.000
3.000 0.000
4.000 0.000
5.000 0.200
6.000 0.167
7.000 0.143
8.000 0.250
9.000 0.222
10.000 0.200
11.000 0.182
12.000 0.250
13.000 0.231
14.000 0.214
15.000 0.200
16.000 0.250
17.000 0.235
18.000 0.222
19.000 0.263
20.000 0.250
21.000 0.238
22.000 0.273
23.000 0.261
24.000 0.250
25.000 0.240
26.000 0.231
27.000 0.222
28.000 0.214
29.000 0.207
30.000 0.233
31.000 0.226
32.000 0.219
33.000 0.212
34.000 0.235
35.000 0.229
36.000 0.222
37.000 0.216
38.000 0.211
39.000 0.205
40.000 0.200
41.000 0.195
42.000 0.190
43.000 0.186
44.000 0.182
45.000 0.178
46.000 0.174
47.000 0.170
48.000 0.167
49.000 0.163
50.000 0.160
51.000 0.157
52.000 0.154
53.000 0.151
54.000 0.148
55.000 0.145
56.000 0.143
57.000 0.140
58.000 0.138
59.000 0.136
60.000 0.133
61.000 0.131
62.000 0.129
63.000 0.127
64.000 0.125
65.000 0.123
66.000 0.121
67.000 0.119
68.000 0.118
69.000 0.116
70.000 0.114
71.000 0.113
72.000 0.111
73.000 0.110
74.000 0.108
75.000 0.107
76.000 0.105
77.000 0.104
78.000 0.103
79.000 0.101
80.000 0.100
|
File: _classplot_dat2
# GNUPLOT data file for Uncertain hits, series 2
1.000 0.000
2.000 0.000
3.000 0.000
4.000 0.000
5.000 0.000
6.000 0.000
7.000 0.000
8.000 0.000
9.000 0.000
10.000 0.000
11.000 0.000
12.000 0.000
13.000 0.000
14.000 0.000
15.000 0.000
16.000 0.000
17.000 0.000
18.000 0.000
19.000 0.000
20.000 0.000
21.000 0.000
22.000 0.000
23.000 0.000
24.000 0.000
25.000 0.000
26.000 0.038
27.000 0.037
28.000 0.036
29.000 0.034
30.000 0.033
31.000 0.032
32.000 0.031
33.000 0.030
34.000 0.029
35.000 0.029
36.000 0.028
37.000 0.054
38.000 0.053
39.000 0.051
40.000 0.050
41.000 0.049
42.000 0.048
43.000 0.047
44.000 0.045
45.000 0.044
46.000 0.043
47.000 0.064
48.000 0.062
49.000 0.061
50.000 0.060
51.000 0.059
52.000 0.058
53.000 0.057
54.000 0.056
55.000 0.055
56.000 0.054
57.000 0.070
58.000 0.069
59.000 0.068
60.000 0.067
61.000 0.066
62.000 0.065
63.000 0.063
64.000 0.062
65.000 0.062
66.000 0.061
67.000 0.060
68.000 0.059
69.000 0.058
70.000 0.057
71.000 0.056
72.000 0.056
73.000 0.055
74.000 0.054
75.000 0.053
76.000 0.053
77.000 0.052
78.000 0.051
79.000 0.051
80.000 0.050
|
File: _classplot_dat3
# GNUPLOT data file for Unknown hits, series 3
1.000 0.000
2.000 0.000
3.000 0.000
4.000 0.000
5.000 0.000
6.000 0.000
7.000 0.000
8.000 0.000
9.000 0.000
10.000 0.000
11.000 0.000
12.000 0.000
13.000 0.000
14.000 0.000
15.000 0.000
16.000 0.000
17.000 0.000
18.000 0.000
19.000 0.000
20.000 0.000
21.000 0.000
22.000 0.000
23.000 0.000
24.000 0.000
25.000 0.000
26.000 0.000
27.000 0.000
28.000 0.000
29.000 0.000
30.000 0.000
31.000 0.000
32.000 0.000
33.000 0.000
34.000 0.000
35.000 0.000
36.000 0.000
37.000 0.000
38.000 0.026
39.000 0.026
40.000 0.025
41.000 0.049
42.000 0.048
43.000 0.047
44.000 0.068
45.000 0.067
46.000 0.065
47.000 0.064
48.000 0.083
49.000 0.082
50.000 0.080
51.000 0.098
52.000 0.096
53.000 0.094
54.000 0.111
55.000 0.109
56.000 0.107
57.000 0.105
58.000 0.121
59.000 0.119
60.000 0.117
61.000 0.131
62.000 0.129
63.000 0.127
64.000 0.125
65.000 0.123
66.000 0.121
67.000 0.119
68.000 0.118
69.000 0.116
70.000 0.114
71.000 0.113
72.000 0.111
73.000 0.110
74.000 0.108
75.000 0.107
76.000 0.105
77.000 0.104
78.000 0.103
79.000 0.101
80.000 0.100
|
File: _classplot_dat4
# GNUPLOT data file for False hits, series 4
1.000 0.000
2.000 0.000
3.000 0.000
4.000 0.000
5.000 0.000
6.000 0.000
7.000 0.000
8.000 0.000
9.000 0.000
10.000 0.000
11.000 0.000
12.000 0.000
13.000 0.000
14.000 0.000
15.000 0.000
16.000 0.000
17.000 0.000
18.000 0.000
19.000 0.000
20.000 0.000
21.000 0.000
22.000 0.000
23.000 0.000
24.000 0.042
25.000 0.040
26.000 0.038
27.000 0.037
28.000 0.036
29.000 0.034
30.000 0.033
31.000 0.032
32.000 0.031
33.000 0.030
34.000 0.029
35.000 0.029
36.000 0.028
37.000 0.027
38.000 0.026
39.000 0.051
40.000 0.075
41.000 0.073
42.000 0.095
43.000 0.116
44.000 0.114
45.000 0.111
46.000 0.109
47.000 0.106
48.000 0.104
49.000 0.122
50.000 0.140
51.000 0.137
52.000 0.154
53.000 0.170
54.000 0.167
55.000 0.164
56.000 0.161
57.000 0.158
58.000 0.155
59.000 0.169
60.000 0.183
61.000 0.180
62.000 0.194
63.000 0.206
64.000 0.219
65.000 0.231
66.000 0.242
67.000 0.254
68.000 0.265
69.000 0.275
70.000 0.286
71.000 0.296
72.000 0.306
73.000 0.315
74.000 0.324
75.000 0.333
76.000 0.342
77.000 0.351
78.000 0.359
79.000 0.367
80.000 0.375
|
File: _summary
rocplot summary file (15 Jul 2013)
mode == 2 (Multiple input file mode)
multimode == 2 (Combine data: single ROC plot, single classification plot.)
datamode == 1 (Single list of known true relatives.)
File Known
data1.hits 140
data2.hits 140
ROC50 == 0.185 (combined)
|
File: rocplot.log
MODE INFO
modei: 2
multimodei: 2
datamodei: 1
NUMBER OF INPUT FILES
numfiles: 2
NAMES ONLY OF INPUT FILES
hitsnames[0]: data1.hits
hitsnames[1]: data2.hits
ROC NUMBER
roc: 50
ROC VALUES
rocn[0]: 0.184714
COUNT OF HITS
hitcnt[0]: 80
|
5.0 DATA FILES
ROCPLOT does not use a data file.
6.0 USAGE
6.1 COMMAND LINE ARGUMENTS
Perform ROC analysis on hits files
Version: EMBOSS:6.6.0.0
Standard (Mandatory) qualifiers (* if not always prompted):
[-hitsfilespath] dirlist [rocplot] This option specifies the
directory of hits files (input). A 'hits
file' contains a list of hits (e.g. from a
prediction method) that are classified and
rank-ordered on the basis of score, p-value,
E-value etc. The files generated by using
SIGSCAN and LIBSCAN will contain the results
of a search of a discriminating element
(e.g. hidden Markov model, profile or
signature) against a sequence database. The
ROCPLOT application is run on the files to
perform Receiver Operator Characteristic
(ROC) analysis on the hits.
-mode menu [1] This option specifies the mode of
ROCPLOT operation (main mode). In 'single
input file mode', ROC analysis is performed
on the individual hits file; a ROC plot
containing a single ROC curve, and a single
ROC value and classification plot are
generated. In 'multiple input file mode'
there are two sub-modes depending upon
whether (1) ROC analysis is to performed
separately for the individual input files or
(2) the lists of hits in the hits files are
combined and ROC analysis is performed on
the whole (see the ACD option called
'multimode' for more information). If the
input file does not contain at least as many
'FALSE' hits as are specified after the
'ROC' token in the input file, then an error
will be generated and rocplot will
terminate. Where multiple input files are
given as input, each must contain the same
value after the 'ROC' token, or an error
will be generated and rocplot will
terminate. The hits in the hits files *must*
have been rank-ordered on the basis of
score, p-value, E-value etc, with the
highest scoring / most significant hit being
given in the highest rank (1); i.e. on the
second line of the file. Other hits should
then be given in order of decreasing score /
significance. (Values: 1 (Single input file
mode); 2 (Multiple input file mode))
* -multimode menu [1] This option specifies the mode of
ROCPLOT operation (multimode). In 'Do not
combine data' mode, ROC analysis is
performed separately for the individual
input files. Multiple ROC curves will be
given on the same ROC plot and a ROC value
and a classification plot will be generated
for each input file. A bar chart giving the
distribution of ROCn values, and the mean
and standard deviation of ROCn values are
also generated. In 'Combine data' mode, the
lists of hits in the hits files are combined
and ROC analysis is performed on the whole.
A single ROC curve will be given in the ROC
plot and a single ROC value and
classification plot will be generated. In
this second mode there are two further
sub-modes depending on whether there is (1)
a single list of known true relatives for
the different searches or (2) there is a
different list of known true relatives for
each different search (see the ACD option
called 'datamode' for more information)
(Values: 1 (Do not combine data (multiple
ROC curves in single ROC plot - multiple
classification plots)); 2 (Combine data
(single ROC curve - single classification
plot)))
* -datamode menu [1] This option specifies the mode of
ROCPLOT operation (datamode). This determine
how the ROC number and value are calculated
in cases where there are multiple input
files (lists of hits) and the user has
specified the data are to be combined. See
rocplot.c for more information. (Values: 1
(Single list of known true relatives); 2
(Multiple lists of known true relatives))
* -thresh integer [10] This option specifies the overlap
threshold for hits. In cases where the lists
of hits are to be combined and there is a
single set of relatives, the accession
number (or other database identifier code)
of the hit, and the start and end point
respectively of the hit relative to full
length sequence must be provided in the
lists of hits (see 'Input file format'
below). rocplot ensures that only unique
hits are counted when calculating SENS and
SPEC; two hits are 'unique' if they have (i)
different accesssion numbers or (ii) the
same accession numbers but which do not
overlap by any more than a user-defined
number of residues. The overlap is
determined from the start and end points of
the hit. For example two hits both with the
same accession numbers and with the start
and end points of 1-100 and 91 - 190
respectively are considered to be the same
hit if the overlap threshold is 10 or less.
(Any integer value)
[-outdir] outdir [./] This option specifies the directory
where output files are written.
[-rocbasename] string [_rocplot] This option specifies the base
name of ROC plot file(s) (output). A file of
meta data that contains graphs that
illustrate the diagnostic performance of the
discriminator. rocplot generates Receiver
Operating Characteristic (ROC) curves, that
display graphically the sensitivity and
specificity of discriminating elements, and
accompanying ROC value(s), which are a
convenient numerical measure of the
sensitivity and specificity of a method.
Classification plots, which are a valuable
aid in interpreting the ROC plot and value,
are also generated and, depending upon the
mode rocplot is run in, a plot of the
distribution of ROC values. (Any string)
-outfile outfile [_summary] This option specifies the name of
the summary file (output). A text file
summarising the analysis.
* -barbasename string [_barchart] This option specifies the base
name of bar chart for ROC value distribution
(output). A bar chart giving the
distribution of ROCn values will be
generated when multiple input files (lists
of hits) are provided and the user has
specified 'Do not combine data (multiple ROC
curves). (Any string)
-classbasename string [_classplot] This option specifies the base
name of classification plot file(s)
(output). Classification plots are a
valuable aid in interpreting the ROC plot
and value. A single plot will be generated
where a single input file is provided or
where multiple input files are provided and
the user has specified 'Combine data (single
ROC curve)' mode. Multiple plots will be
generated where multiple input files are
provided and the user has specified 'Do not
combine data (multiple ROC curves)' mode.
(Any string)
Additional (Optional) qualifiers: (none)
Advanced (Unprompted) qualifiers:
-norange boolean [N] This option specifies whether to
disregard range data when identifying unique
hits. If set, the range data specified in
the hits files are disregarded, two hits are
classed as unique if they have different
accession numbers (no requirement for
overlapping ranges).
-logfile outfile [rocplot.log] Domainatrix log output file
Associated qualifiers:
"-hitsfilespath" associated qualifiers
-extension1 string Default file extension
"-outdir" associated qualifiers
-extension2 string Default file extension
"-outfile" associated qualifiers
-odirectory string Output directory
"-logfile" associated qualifiers
-odirectory string Output directory
General qualifiers:
-auto boolean Turn off prompts
-stdout boolean Write first file to standard output
-filter boolean Read first file from standard input, write
first file to standard output
-options boolean Prompt for standard and additional values
-debug boolean Write debug output to program.dbg
-verbose boolean Report some/full command line options
-help boolean Report command line options and exit. More
information on associated and general
qualifiers can be found with -help -verbose
-warning boolean Report warnings
-error boolean Report errors
-fatal boolean Report fatal errors
-die boolean Report dying program messages
-version boolean Report version number and exit
| Qualifier |
Type |
Description |
Allowed values |
Default |
| Standard (Mandatory) qualifiers |
[-hitsfilespath]
(Parameter 1) |
dirlist |
This option specifies the directory of hits files (input). A 'hits file' contains a list of hits (e.g. from a prediction method) that are classified and rank-ordered on the basis of score, p-value, E-value etc. The files generated by using SIGSCAN and LIBSCAN will contain the results of a search of a discriminating element (e.g. hidden Markov model, profile or signature) against a sequence database. The ROCPLOT application is run on the files to perform Receiver Operator Characteristic (ROC) analysis on the hits. |
Directory with files |
rocplot |
| -mode |
list |
This option specifies the mode of ROCPLOT operation (main mode). In 'single input file mode', ROC analysis is performed on the individual hits file; a ROC plot containing a single ROC curve, and a single ROC value and classification plot are generated. In 'multiple input file mode' there are two sub-modes depending upon whether (1) ROC analysis is to performed separately for the individual input files or (2) the lists of hits in the hits files are combined and ROC analysis is performed on the whole (see the ACD option called 'multimode' for more information). If the input file does not contain at least as many 'FALSE' hits as are specified after the 'ROC' token in the input file, then an error will be generated and rocplot will terminate. Where multiple input files are given as input, each must contain the same value after the 'ROC' token, or an error will be generated and rocplot will terminate. The hits in the hits files *must* have been rank-ordered on the basis of score, p-value, E-value etc, with the highest scoring / most significant hit being given in the highest rank (1); i.e. on the second line of the file. Other hits should then be given in order of decreasing score / significance. |
| 1 | (Single input file mode) | | 2 | (Multiple input file mode) |
|
1 |
| -multimode |
list |
This option specifies the mode of ROCPLOT operation (multimode). In 'Do not combine data' mode, ROC analysis is performed separately for the individual input files. Multiple ROC curves will be given on the same ROC plot and a ROC value and a classification plot will be generated for each input file. A bar chart giving the distribution of ROCn values, and the mean and standard deviation of ROCn values are also generated. In 'Combine data' mode, the lists of hits in the hits files are combined and ROC analysis is performed on the whole. A single ROC curve will be given in the ROC plot and a single ROC value and classification plot will be generated. In this second mode there are two further sub-modes depending on whether there is (1) a single list of known true relatives for the different searches or (2) there is a different list of known true relatives for each different search (see the ACD option called 'datamode' for more information) |
| 1 | (Do not combine data (multiple ROC curves in single ROC plot - multiple classification plots)) | | 2 | (Combine data (single ROC curve - single classification plot)) |
|
1 |
| -datamode |
list |
This option specifies the mode of ROCPLOT operation (datamode). This determine how the ROC number and value are calculated in cases where there are multiple input files (lists of hits) and the user has specified the data are to be combined. See rocplot.c for more information. |
| 1 | (Single list of known true relatives) | | 2 | (Multiple lists of known true relatives) |
|
1 |
| -thresh |
integer |
This option specifies the overlap threshold for hits. In cases where the lists of hits are to be combined and there is a single set of relatives, the accession number (or other database identifier code) of the hit, and the start and end point respectively of the hit relative to full length sequence must be provided in the lists of hits (see 'Input file format' below). rocplot ensures that only unique hits are counted when calculating SENS and SPEC; two hits are 'unique' if they have (i) different accesssion numbers or (ii) the same accession numbers but which do not overlap by any more than a user-defined number of residues. The overlap is determined from the start and end points of the hit. For example two hits both with the same accession numbers and with the start and end points of 1-100 and 91 - 190 respectively are considered to be the same hit if the overlap threshold is 10 or less. |
Any integer value |
10 |
[-outdir]
(Parameter 2) |
outdir |
This option specifies the directory where output files are written. |
Output directory |
./ |
[-rocbasename]
(Parameter 3) |
string |
This option specifies the base name of ROC plot file(s) (output). A file of meta data that contains graphs that illustrate the diagnostic performance of the discriminator. rocplot generates Receiver Operating Characteristic (ROC) curves, that display graphically the sensitivity and specificity of discriminating elements, and accompanying ROC value(s), which are a convenient numerical measure of the sensitivity and specificity of a method. Classification plots, which are a valuable aid in interpreting the ROC plot and value, are also generated and, depending upon the mode rocplot is run in, a plot of the distribution of ROC values. |
Any string |
_rocplot |
| -outfile |
outfile |
This option specifies the name of the summary file (output). A text file summarising the analysis. |
Output file |
_summary |
| -barbasename |
string |
This option specifies the base name of bar chart for ROC value distribution (output). A bar chart giving the distribution of ROCn values will be generated when multiple input files (lists of hits) are provided and the user has specified 'Do not combine data (multiple ROC curves). |
Any string |
_barchart |
| -classbasename |
string |
This option specifies the base name of classification plot file(s) (output). Classification plots are a valuable aid in interpreting the ROC plot and value. A single plot will be generated where a single input file is provided or where multiple input files are provided and the user has specified 'Combine data (single ROC curve)' mode. Multiple plots will be generated where multiple input files are provided and the user has specified 'Do not combine data (multiple ROC curves)' mode. |
Any string |
_classplot |
| Additional (Optional) qualifiers |
| (none) |
| Advanced (Unprompted) qualifiers |
| -norange |
boolean |
This option specifies whether to disregard range data when identifying unique hits. If set, the range data specified in the hits files are disregarded, two hits are classed as unique if they have different accession numbers (no requirement for overlapping ranges). |
Boolean value Yes/No |
No |
| -logfile |
outfile |
Domainatrix log output file |
Output file |
rocplot.log |
| Associated qualifiers |
| "-hitsfilespath" associated dirlist qualifiers
|
-extension1
-extension_hitsfilespath |
string |
Default file extension |
Any string |
|
| "-outdir" associated outdir qualifiers
|
-extension2
-extension_outdir |
string |
Default file extension |
Any string |
|
| "-outfile" associated outfile qualifiers
|
| -odirectory |
string |
Output directory |
Any string |
|
| "-logfile" associated outfile qualifiers
|
| -odirectory |
string |
Output directory |
Any string |
|
| General qualifiers |
| -auto |
boolean |
Turn off prompts |
Boolean value Yes/No |
N |
| -stdout |
boolean |
Write first file to standard output |
Boolean value Yes/No |
N |
| -filter |
boolean |
Read first file from standard input, write first file to standard output |
Boolean value Yes/No |
N |
| -options |
boolean |
Prompt for standard and additional values |
Boolean value Yes/No |
N |
| -debug |
boolean |
Write debug output to program.dbg |
Boolean value Yes/No |
N |
| -verbose |
boolean |
Report some/full command line options |
Boolean value Yes/No |
Y |
| -help |
boolean |
Report command line options and exit. More information on associated and general qualifiers can be found with -help -verbose |
Boolean value Yes/No |
N |
| -warning |
boolean |
Report warnings |
Boolean value Yes/No |
Y |
| -error |
boolean |
Report errors |
Boolean value Yes/No |
Y |
| -fatal |
boolean |
Report fatal errors |
Boolean value Yes/No |
Y |
| -die |
boolean |
Report dying program messages |
Boolean value Yes/No |
Y |
| -version |
boolean |
Report version number and exit |
Boolean value Yes/No |
N |
6.2 EXAMPLE SESSION
An example of interactive use of ROCPLOT is shown below.
Here is a sample session with rocplot
% rocplot
Perform ROC analysis on hits files
Hits directories [rocplot]: rocplot/hitsin
Available modes
1 : Single input file mode
2 : Multiple input file mode
Select mode of operation. [1]: 2
Available modes
1 : Do not combine data (multiple ROC curves in single ROC plot - multiple classification plots)
2 : Combine data (single ROC curve - single classification plot)
Select mode of operation. [1]: 2
Available modes
1 : Single list of known true relatives
2 : Multiple lists of known true relatives
Select mode of operation. [1]: 1
Overlap threshold for hits. [10]:
General output file output directory [./]:
Base name of ROC plot file(s) (output). [_rocplot]:
Rocplot summary output file [_summary]:
Base name of classification plot file(s) (output). [_classplot]:
/homes/user/test/data/structure/rocplot/hitsin/data1.hits
/homes/user/test/data/structure/rocplot/hitsin/data2.hits
Processing data1.hits
Processing data2.hits
Please wait ... done!
|
Go to the output files for this example
7.0 KNOWN BUGS & WARNINGS
GNUPLOT must be started in the same directory as the gnuplot data files.
If you run ROCPLOT on many input files without specifying combination of
data the ROC plot generated can get very cluttered. This is not a flaw of
ROCPLOT, but an inevitable consequence of trying to draw too many things
on the same plot. The recomended maximum is 5 to 10 input files.
The hits in the hits files *must* be rank-ordered on the basis of score,
p-value, E-value etc, with the highest scoring / most significant hit given
in the highest rank (1); i.e. on the second line of the file. Other hits
should then be given in order of decreasing score / significance.
8.0 NOTES
Future implementation
1. Accept a feature file as input.
2. Split ROCPLOT into separate programs, one for each of the major modes.
Description of 'sort' mode (additional option in ACD)
This option specifies whether to process the input files in blocks (of the same domain identifier). In this case the analysis mode (mode-multimode-datamode) are set to Multiple input file - combine data - Single list of known true relatives (2-2-1) and the analysis is performed on each block of hits files with the same domain identifier. In the output file, ROC values are given for each combined analysis and the mean and SD of all the combined analyses are given. The domain identifier is defined as the text between the first and second period ('/.') in the input file name.
Description of 'norange' mode (additional option in ACD)
This option specifies whether to disregard range data when identifying unique hits. If set, the range data specified in the hits files are disregarded, two hits are classed as unique if they have different accession numbers (no requirement for overlapping ranges).
8.1 GLOSSARY OF FILE TYPES
| FILE TYPE |
FORMAT |
DESCRIPTION |
CREATED BY |
SEE ALSO |
| Hits file |
Text file of classified hits |
A list of hits (e.g. from a prediction method) that are classified and rank-ordered on the basis of score, p-value, E-value etc. |
ROCON and LIBSCAN (hits from searches of a discriminating element (hidden Markov model, profile or signature) against a sequence database). |
ROCPLOT is run on the files to perform Receiver Operator Characteristic (ROC) analysis on the hits. |
None
9.0 DESCRIPTION
Predictive methods are a mainstay of bioinformatics. Discrciminating
elements such as hidden Markov models (HMM), sparse protein signatures
and profiles can be generated for a set of proteins with related sequence,
structural or functional properties. These discriminators are
characteristic of the property considered and can be used diagnostically,
for instance, by screening a database of uncharacterised sequences. When
assessing predictive performance a "gold standard" of truth is required.
This is a set of examples that are known to be related to the discriminating
element, and, ideally, a further set that is known to be definitely not
related. For example, to assess a protein family HMM to detect true members
of that family requires, at least, a list of the known family members. If a
method works well for the "gold standard" we can infer it will work well
generally. Traditionally, swissprot annotation was used but this is somewhat
unreliable because the annotation is derived from sequence comparison as well
as experimental data. Increasingly, use is made of databases such as SCOP,
in which sequence, structural and functional relationships are classified.
As an aside, such databases are biased for domains, which are the unit of
classification, so it's important to check that a method tested on e.g. SCOP
will also work on full-length sequences.
9.1 Sensitivity and specificity
Most predictive methods can be placed into two broad groupings: (i) Methods
that produce a definite yes/no answer. There is a single list of "hits" and
things not in this list are "misses". (ii) Methods that produce a list of
hits that is rank-ordered on the basis of the score or p-value of the
discrimintor-sequence match. The hit with the highest / most significant
score will be in highest rank, i.e. rank 1. Usually, a cutoff value of rank,
score or p-value is applied; "hits" occur at and above the cuttoff and
"misses" occur below it.
Armed with the notion of a "gold standard" and "hits" and "misses", all hits
retrieved by a search can be organised as in Figure 4.
Figure 4 Classification of hits
From the gold standard
| | |
| Related | Unrelated|
_______|__________|__________|_______
| | |
S r (+ve) | TP | FP | P (=TP+FP)
e e hits | | |
a s _______|_ ________|__________|_______
r u | | |
c l (-ve) | FN | TN | N (=FN+TN)
h t misses | | |
_______|__________|__________|_______
| | |
| R | U |
| (=TP+FN) | (=FP+TN) |
|
Where TP are true positives, FN are false negatives, R (TP+FN) is the total
number of known true hits (relatives). FP are false positives, TN are true
negatives and U (FP+TN) is the total number of known non-relatives. The
number of positives is given by P (TP+FP) and the number of misses by N
(FN+TN).
The two basic types of error are where (i) a relationship is missed ("false
negative" or "ommission error") and (ii) a relationship is inferred which
does not truly exist ("false positive" or "commission error"). The cost of
these two errors are not usually equal: it depends on the specific
application but usually false positives are worse than false negatives. A
crude way to measure the performance is to quote ommission and commission
error rates at a fixed cutoff value to the list of hits. These rates are
usually given as sensitivity (SENS or "coverage") and specificity (SPEC or
"accuracy") of the method and are defined as follows.
SENS = TP / R
SPEC = TP / P
Another measure of specificity (JMB 282, 903-918) defines SENS = TN / U. The
measure used depends on the specific application, but TP/P is often most
suitable as it reflects the hits that are actually retrieved by the search.
TP / P is used in ROCPLOT (see Section 10.2).
The most basic graphical representation of sensitivity and specificity is
the "coverage versus error plot" or "sensitivity curve" (Figure 5). This
plots the number of true positives detected (y-axis) versus the number of
false positives detected (x-axis), at different cutoff values in the list of
hits. The word 'detected' here refers to a hit that is above the cutoff,
i.e. is of a higher or more significant score.
Figure 5 A "coverage versus error" plot
|
| *
No. true | *
positives | *
detected | *
| *
| *
| *
| *
| *
|*
|______________________________
No. false
positives
detected
|
9.2 ROC plot
A superior measure of diagnostic performance is to use Receiver Operator
Characteristic (ROC) curves to display graphically the sensitivity and
specificity of a method. ROC analysis is a powerful aid to interpretation
and has been widely used, for instance to evaluate clinical diagnostic tests
and in the bioinformatics literature. A ROC curve (Figure 6) is a
generalised version of the "coverage versus error" plot. It plots SENS
(TP/R) on the y-axis, i.e. the fraction of known true hits detected or the
"rate of true positives", versus 1-SPEC (1 - TP/P) on the x-axis, i.e. 1
minus the fraction of detected hits that are true positives or the "rate of
false positives". ROC curves are generated by plotting SENS versus (1-SPEC)
for all possible cutoff values in a rank-ordered list of hits.
Figure 6 A ROC curve
|
| *
SENS | *
| *
"rate of | *
true | *
positives" | *
| *
| *
| *
|*
|______________________________
1 - SPEC
"rate of false positives"
|
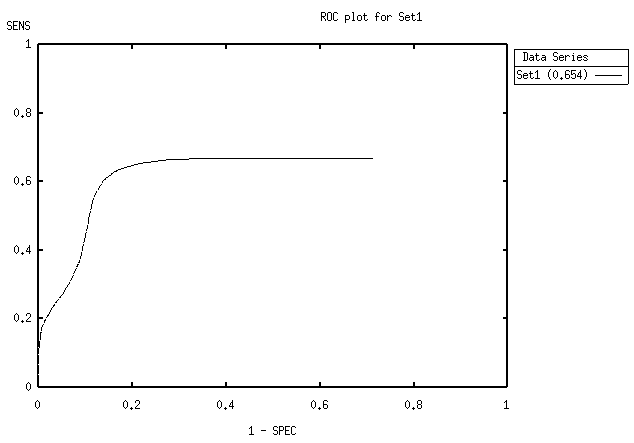
The first image is a schematic, the second is a screenshot of the a ROCPLOT-generated roc plot, visualised by using GNUPLOT.
A ROC curve shows the trade-off between sensitivitiy and specificity: as
sensitivitiy increases, specificity decreases. The ideal ROC curve lies on
the y-axis, i.e. there is perfect discrimination between related and
unrelated proteins. A ROC curve for a good prediction should always be to
the left of the diagonal. ROC curves are very useful for comparing two
diffent methods (e.g. homology search methods) because if one method produces
a curve to the left of another then that method is superior, regardless of
the cost of ommission and commission errors.
9.3 ROC value
The area under the ROC curve (AUROC) gives the probability of a correct
classification and is a very convenient numerical measure of the sensitivity
and specificity of a method. Areas are relative to a ROC space which is a
unit square in which both SENS and SPEC are plotted from 0 to 1. An area of
0.9 for example means that a sequence from the group of known relatives has
a probability of 0.9 of scoring higher than a sequence from the group of
known non-relatives. The best possible prediction has an AUROC of 1.
In most cases however there are vastly more true negatives than true
positives. This is the case when a search is made with a sequence against a
large sequence database. As most sequence are quite discriminating for
their family, the AUROC for a ROC curve plotted for the results of the entire
database search will be very close to 1. The AUROC value is still useful but
it has to be calculated to 5 or 6 decimal places. Furthermore all the curves
would look identical which makes comparing two methods by eye impossible, all
the database scores would have to be written to disk, and the value does not
really represent the way in which the average biologist, who is unprepared to
inspect many thousands of false positives, would use the method. For these
reasons, ROC curves are usually truncated to the first 50 or 100 false hits,
and the so-called ROC50 or ROC100 value calculated. ROCn values are quicker
and more convenient to calculate, can be expressed by fewer decimal places
and reflect the way in which the average biologist will use the method.
9.4 Classification plot
In many cases not every hit returned by a search can be clearly classified as
true or false or it might otherwise be desirable to manage hits with an
intermediate classification. This might be the case where the gold standard
is based on a hierarchic structure (e.g. SCOP). Consider conceptual "cross",
"uncertain" and "unknown" hits. "Cross hits" have a definite relation to the
query but not at such a fine level as a "true" hit. An example is a query
matching a sequence belonging to a different family but the same superfamily
as the query. An "uncertain hit" might show some but not clear evidence of a
relation. An example would be a query matching a sequence belonging to a
different family and superfamily, but the same fold as the query. For other
hits, nothing may be known either way and these would be classified as
"unknown". ROCPLOT supports "cross", "uncertain" and "unknown" hits and
provides a graphical representation of the classifications of hits by
generating a "classification plot".
A classification plot (Figure 7) shows the proportion of hits detected that
are 'true', 'cross', 'uncertain', 'unknown' and 'false'. The y-axis is the
proportion of the hits detected that are of a certain type, the x-axis is
the proportion of the total number of hits detected. A separate curve is
given for hits of each type. In ROCPLOT a classification plot is generated
by plotting these proportions at each rank in the list of hits up to the
point where a user-defined number of 'false' hits are detected. As ROC plots
and values (see below) do not consider 'cross', 'uncertain' and 'unknown'
hits, the classification plot is a useful aid in interpreting the ROC plot
and value for some applications.
Figure 7 A classification plot
Proportion of 1.0|
hits detected |
that are of a |
certain type |
| * * TRUE
| * . . CROSS
| * .
| * .
| * . x x FALSE
| *. x
|*. x
|______________________________
0 1.0
Proportion of total
number of hits detected.
|
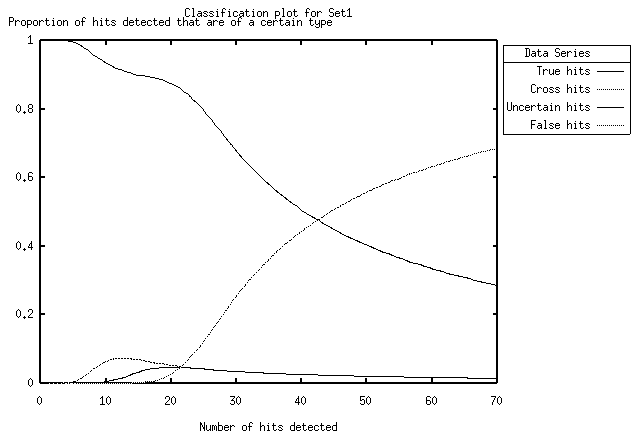
The first image is a schematic (hits of classification 'uncertain' and 'unknown' are not shown for clarity). The second is an screenshot of the a ROCPLOT-generated classification plot, visualised by using GNUPLOT.
9.5 Processing multiple lists of hits (no combination of lists)
ROC analysis is a powerful way to compare predictive methods side by side.
A ROC value can be generated for each method and a curve plotted on the same
ROC plot. For some applications a summary of a set of ROC values is required.
Depending upon mode (see Section 2.1), ROCPLOT will generate the mean,
standard deviation (SD) and a bar chart (Figure 8) of the distribution of
ROCn values. In constructing the bar chart, the range of possible ROC values
from 0 to 1 is divided into 20 bins of size 0.05 and the frequency of
occurence of ROC values in each bin range is calculated.
Figure 8 Bar chart for distribution of ROCn values
Frequency |
| ___
| | |
| ___| |
| ___ | | |
| | | | | |
| ___ | | | | |
| | |___| | | | |
| | | | |___| | |
|___| | | | | | |
| | | | | | | |
|___|___|___|___|___|___|___|__
Bins for different
ranges of value of
ROCn value
|
9.6 Processing multiple lists of hits (combination of lists)
In some cases it is desirable to combine data from multiple lists of hits and
derive a single ROC curve and value. Such cases fall into one of two broad
groups: (i) There is a single set of known true relatives for the different
searches, for example, when assessing the performance of multiple
discriminating elements for a single family. In these cases the typical
ROC50 or ROC100 value is generated. (ii) There is a different set of known
true relatives for each different search, for example, when assessing the
performance of a single discriminating element over mutliple families. A
much higher ROC number is used. For exmaple, ROC500 is reasonable if 10
lists of hits are combined.
Lists of hits arising from different searches can be combined and reordered
if they are scored on the same scoring scale or have been assigned a p-value.
In principle one way to use ROCPLOT is to do the combination and reordering
yourself and provide ROCPLOT with a single list of hits as input. This,
however, is not possible if the lists of hits use different scoring schemes
and a p-value is not available. Furthermore, in many cases the relative
positioning of hits in the list is more important than the absolute score.
If two lists of hits (A and B) whose hits lie on different regions of the
same scoring scale are merged and reordered, true hits, which rank very
highly in their own list (A), might be relegated way down the merged list,
appearing after false hits from list B. Therefore the high-ranking and
potentially interesting hits in list A might, depending on the ROCn value
calculated, not be considered in the combination ROC value. To overcome
this, the lists of hits can be processed in parallel: to consider all the
hits at rank 1 in the different lists first, then all the hits at rank 1
and 2, and so on. This is the approach taken in ROCPLOT (see Section 10).
10.0 ALGORITHM
10.1 Classification plot
The proportion of the total hits detected that are of a certain type (TRUE,
CROSS, UNCERTAIN, UNKNOWN and FALSE) is calculated at each rank position in
the list of hits, from the first rank (hit) up to and including the hit
corresponding to the nth false positive. n is the ROC number given in the
hits file. For example, if i is the current rank number,
Proportion(TRUE) = (Number of TRUE tokens from ranks 1 to i / i).
10.2 ROC plot
10.2.1 "Single hits file" mode and "Multiple hits files - Do not
combine data" mode
SENS and SPEC are calculated at each rank in the list of hits from the first
rank up to and including the hit that is the nth false positive. n is the
ROC number given in the hits file. SENS and SPEC are calculated as follows.
SENS(i) = TP / R
SPEC(i) = TP / i
Where i is the current rank number, TP is the number of TRUE tokens occuring
from rank 1 to i. R is the total number of known true hits (relatives)
specified after the 'RELATED' token in the hits file(s) (see Section 3.1).
Hits classified as CROSS, UNCERTAIN and UNKNOWN are all treated as FALSE.
This means that the ROC curve is really giving "rate of noise" on the x-axis
rather than the "rate of false positives". The "noise" might actually
include genuinely interesting hits and for this reason, the ROC plot must be
interpreted in the light of the classification plot if CROSS, UNCERTAIN and
UNKNOWN classifications are used. If the hits file contains fewer than n
hits that are non-TRUE, an error is generated and ROCPLOT terminates.
10.2.2 "Multiple hits files" / "Combine data" mode
SENS and SPEC are calculated at different ranks as before but this time the
lists are processed in parallel. SENS and SPEC are calculated from each list
in turn at each rank from the first rank up to and including the rank at
which n false positive (from the different lists) are detected. If there are
5 hits files for example, a maximum of 5 hits are considered to yield up to 5
SENS and 5 SPEC values at each rank. In "Single gold standard" mode, n is
the ROC number specified after the 'ROC' token in the hits files. In
"Multiple gold standard" mode, n = (ROC number from hits files * number of
input files). SENS and SPEC are calculated as follows.
SENS(i, j) = TP / R
SPEC(i, j) = TP / nhits
Where i is the current rank number and j is the number of the list of the hit
being considered. TP is the number of true positives. TP = (Number of TRUE
tokens in ranks 1 to i-1 in all lists + number of TRUE tokens in rank i in
lists 1 to j). Note that in "Single gold standard" mode only those TRUE
tokens corresponding to unique hits (see below) are counted. R is the number
of known 'true' hits (relatives). In "Single gold standard" mode, R equals
the value after the 'RELATED' token in the hits files. In "Multiple gold
standard" mode, R equals the sum of the values given after the 'RELATED'
tokens. nhits is the number of hits considered so far. If the hits files
contain equal numbers of hits, nhits = (i-1)*N + j, where N is the total
number of hits files.
10.3 ROC value
10.3.1 "Single hits file" mode and "Multiple hits files - Do not combine
data" mode
The ROCn value is defined as:
ROCn = 1/nR * T (T is Ti summed for 1<=i<=n)
n is the ROC number from the hits file. R is the total number of known true
hits given in the hits file after the 'RELATED' token. Ti is the number of
TRUE tokens occuring from rank 1 up to the rank for the ith non-TRUE hit.
In other words, Ti is the number of 'true' hits detected above the ith 'false'
hit.
10.3.2 "Multiple hits files" / "Combine data" mode
Again, the ROCn value is defined as :
ROCn = 1/nR * T (T is Ti summed for 1<=i<=n)
n is the ROC number used. In "Single gold standard" mode, n is the ROC
number given in the hits files. In "Multiple gold standard" mode, n = (ROC
number given in hits files * number of input files). R is the number of
known true hits (relatives). In "Single gold standard" mode, R equals the
value given after the 'RELATED' token in the hits files. In "Multiple gold
standard" mode, R equals the sum of the values given after the 'RELATED'
tokens.
Ti is the number of TRUE tokens found up to the ith token that is not 'TRUE'.
If k and j are the rank and number of list respectively at which the nth
non-TRUE hit is detected, Ti = (number of TRUE tokens in ranks 1 to k-1 in
all lists + number of TRUEn tokens in rank k in lists 1 to j). Again, Ti
is the number of 'true' hits detected above the ith 'false' hit.
10.4 Identifying unique hits
In "Multiple hits files" - "Combine data" - "Single gold standard" mode,
ROCPLOT only counts unique hits when calculating SENS and SPEC. Two hits
are 'unique' if they have (i) different accesssion numbers or (ii) the same
accession numbers but which do not overlap by any more than a user-defined
number of residues. The overlap is determined from the start and end points
of the hit. For example two hits, with the same accession numbers and start
and end points of 1-100 and 91 - 190 respectively, are not unique if the
overlap threshold is 10 or less. Duplicate hits (the second and subsequent
occurences of non-unique ones) in the hits files are discarded - they are
NOT considered when calculating the ROC curve and value.
The different hits files might contain different numbers of hits and
therefore at higher ranks, SENS and SPEC might only consider hits from a
subset of all the hits files, up to the last rank for which it is likely
just a single hit will be considered. This is illustrated in Figure 9,
which shows the lists of hits for 3 hits files, a ROC number of 3 is given
for each one. At ranks 1 up to 6, SENS and SPEC would consider hits from
all 3 input files. At rank 7 however, only hits from files 2 and 3 would
be considered as 3 false hits have been detected in file 1 and no more hits
are listed. Similarly at ranks 10 and 11 only hits from file 3 will be
considered.
Figure 9 Calculation of ROC value for multiple hits files
Rank File1 File2 File3
ROC3 ROC3 ROC3
1 TRUE TRUE TRUE
2 TRUE TRUE TRUE
3 TRUE TRUE TRUE
4 FALSE TRUE TRUE
5 FALSE TRUE TRUE
6 FALSE FALSE TRUE
7 FALSE FALSE
8 TRUE FALSE
9 FALSE TRUE
10 TRUE
11 FALSE
|
11.0 RELATED APPLICATIONS
| Program name |
Description |
|---|
| cathparse |
Generate DCF file from raw CATH files |
| domainalign |
Generate alignments (DAF file) for nodes in a DCF file |
| domainnr |
Remove redundant domains from a DCF file |
| domainrep |
Reorder DCF file to identify representative structures |
| domainseqs |
Add sequence records to a DCF file |
| domainsse |
Add secondary structure records to a DCF file |
| helixturnhelix |
Identify nucleic acid-binding motifs in protein sequences |
| libgen |
Generate discriminating elements from alignments |
| matgen3d |
Generate a 3D-1D scoring matrix from CCF files |
| pepcoil |
Predict coiled coil regions in protein sequences |
| rocon |
Generate a hits file from comparing two DHF files |
| scopparse |
Generate DCF file from raw SCOP files |
| seqalign |
Extend alignments (DAF file) with sequences (DHF file) |
| seqfraggle |
Remove fragment sequences from DHF files |
| seqsort |
Remove ambiguous classified sequences from DHF files |
| seqwords |
Generate DHF files from keyword search of UniProt |
| ssematch |
Search a DCF file for secondary structure matches |
12.0 DIAGNOSTIC ERROR MESSAGES
For purposes of generating the ROC plot and ROC curve, hits classified as
CROSS, UNCERTAIN and UNKNOWN are all treated as FALSE. An error is
generated and ROCPLOT terminates in the following cases.
If the hits file contains more TRUE hits than the number after the
'RELATED' token.
In "Multiple hits files" mode, if different values are given after the
'ROC' token in the files.
The number of non-TRUE hits is less than the value after the 'ROC' token.
In "Single gold standard" mode, if different values are given after the
'RELATED' token in the files.
13.0 AUTHORS
Jon Ison (jison@ebi.ac.uk)
The European Bioinformatics Institute
Wellcome Trust Genome Campus
Cambridge CB10 1SD
UK
14.0 REFERENCES
Please cite the authors and EMBOSS.
Rice P, Longden I and Bleasby A (2000) "EMBOSS - The European
Molecular Biology Open Software Suite" Trends in Genetics,
15:276-278.
See also http://emboss.sourceforge.net/
14.1 Other useful references
Gribskov M, Robinson NL. 1996. Use of Receiver Operating Characteristic (ROC) Analysis to Evaluate Sequence Matching. Computers & Chemistry 20(1): 25-33.

